Research
By integrating chemistry, biology and artificial intelligence, the Yang laboratory is developing novel biocatalytic strategies to tackle daunting challenges in synthesis, catalysis, and biomolecular design and engineering. First, we are addressing long-standing problems in asymmetric catalysis by taking advantage of the intimate enzyme-substrate interactions that can be easily tuned via directed evolution. Second, by creatively coupling small-molecule catalysis and biocatalysis, we are advancing novel enzymatic processes which are both new-to-biology and new-to-chemistry. Third, we are developing artificial intelligence and laboratory automation tools to accelerate the de novo design and development of customized biocatalysts and to understand enzyme structure-activity relationships. Using an interdisciplinary approach combining synthetic chemistry, synthetic biology, computational chemistry and computational biology, we reprogram and invent biosynthetic machineries to access reaction space well beyond the native biochemical landscape. Collectively, these efforts will yield fundamental insights into enzymology and chemical reactivity, enabling the rapid assembly of small molecules and macromolecules with biomedical and materials relevance.
Metalloredox Radical Biocatalysis
40-50% natural proteins are metalloproteins. By leveraging the unique redox property of first-row transition-metal cofactors spanning a wide potential window, we repurpose and evolve natural metalloenzymes to catalyze unnatural free radical reactions in a stereocontrolled fashion. Our group is the first to coin and implement the general concept of “metalloredox radical biocatalysis” to impose stereocontrol over fleeting radical intermediates. Using metalloredox radical biocatalysis, we advanced a collection of atom transfer-type free radical reactions, whose asymmetric catalysis has long eluded synthetic organic chemists.
Further reading:
“Stereodivergent atom-transfer radical cyclization by engineered cytochromes P450“ Qi Zhou, Michael Chin, Yue Fu, Peng Liu, and Yang Yang*, Science 2021, 374, 1612–1616.
“Biocatalytic enantioselective C(sp3)–H fluorination enabled by directed evolution of nonheme Fe enzymes“ Liu-Peng Zhao, Binh Khanh Mai, Lida Cheng, Fangqiu Gao, Yunlong Zhao, Rui Guo, Hao Wu, Yongda Zhang, Peng Liu* and Yang Yang*, Nature Synthesis 2024, 3, 967–975.
“Cooperative Photometallobiocatalysis: Nonheme Fe Enzyme-Catalyzed Enantioconvergent Radical Decarboxylative Azidation, Thiocyanation and Isocyanation of Redox-Active Esters” Liu-Peng Zhao, Ken Lin, Pei-Pei Xie, Huichong Liu, Hengye Xiang, Xin Liu, Yunlong Zhao, Peng Liu* and Yang Yang*, Angewandte Chemie International Edition 2025, e202506361.
Pyridoxal Radical Biocatalysis
Pyridoxal 5’-phosphate (PLP) enzymes are powerful biocatalysts responsible for the construction and destruction of amino acids in natural metabolism. By converting natural two-electron PLP enzymes to catalyze single electron processes, we develop pyridoxal radical biocatalysis to enable the stereoselective synthesis of a range of valuable non-canonical amino acids (ncAAs), including those possessing multiple contiguous stereocenters that are difficult to prepare by other means. Our group is the first to design and implement “synergistic photoredox-pyridoxal radical biocatalysis” as a general strategy for convergent ncAA synthesis without protecting groups through a radical mechanism.
Further reading:
“Stereoselective Amino Acid Synthesis by Synergistic Photoredox-Pyridoxal Radical Biocatalysis” Lei Cheng, Dian Li, Binh Khanh Mai, Zhiyu Bo, Lida Cheng, Peng Liu* and Yang Yang*, Science 2023, 381, 444–451.
“Stereoselective amino acid synthesis by photobiocatalytic oxidative coupling“ Tian-Ci Wang, Binh Khanh Mai, Zheng Zhang, Zhiyu Bo, Jiedong Li, Peng Liu* and Yang Yang*, Nature 2024, 629, 98–104.
“Diversity-Oriented Photobiocatalytic Synthesis via Stereoselective Three-Component Radical Coupling” Chen Zhang†, Jun Zhou†, Pei-Pei Xie, Silvia M. Rivera, Turki M. Alturaifi, James Finnigan, Simon Charnock, Peng Liu, and Yang Yang*, Science 2025, First Release, DOI: 10.1126/science.adx2935.
Machine Learning-Guided Enzyme Design and Enzyme Engineering
We apply the state-of-the-art machine learning (ML) methods to de novo enzyme design. Using our methods, guided by careful chemical and structural considerations, we generate novel enzymes to address ongoing challenges in asymmetric catalysis and biochemistry. Integrating sequence, structural and evolutionary information of enzymes, we are also developing next-generation machine learning-guided directed evolution to speed up enzyme engineering.
Further reading:
“Machine learning-guided co-optimization of fitness and diversity facilitates combinatorial library design in enzyme engineering” Kerr Ding†, Michael Chin†, Yunlong Zhao†, Binh Khanh Mai, Wei Huang, Huanan Wang, Peng Liu, Yang Yang* and Yunan Luo*, Nature Communications 2024, 15, 6392.
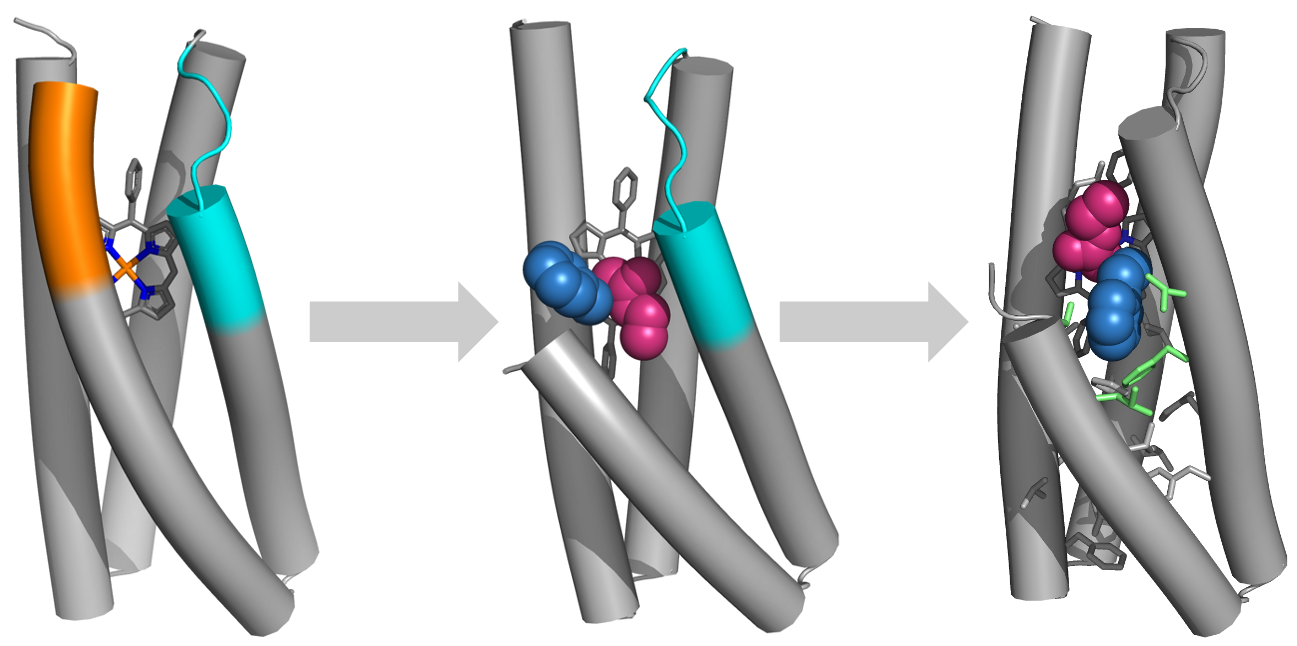
“De Novo Design of Porphyrin-Containing Proteins as Efficient and Stereoselective Catalysts“ Kaipeng Hou†, Wei Huang†, Miao Qi, Thomas H. Tugwell, Turki M. Alturaifi, Yuda Chen, Xingjie Zhang, Lei Lu, Samuel I. Mann, Peng Liu, Yang Yang* and William F. DeGrado*, Science 2025, 388, 665-670.
New Materials at the Chemistry-Biology Interface
In addition to our focus on catalysis at the chemistry-biology interface, we are broadly interested in materials science, particularly polymer chemistry and protein materials. Current focus lies in the directed evolution of novel enzymes for polymer and materials chemistry. Combining our expertise of synthetic chemistry, protein engineering and materials chemistry, we are also pursuing the study of protein materials and engineered living materials.